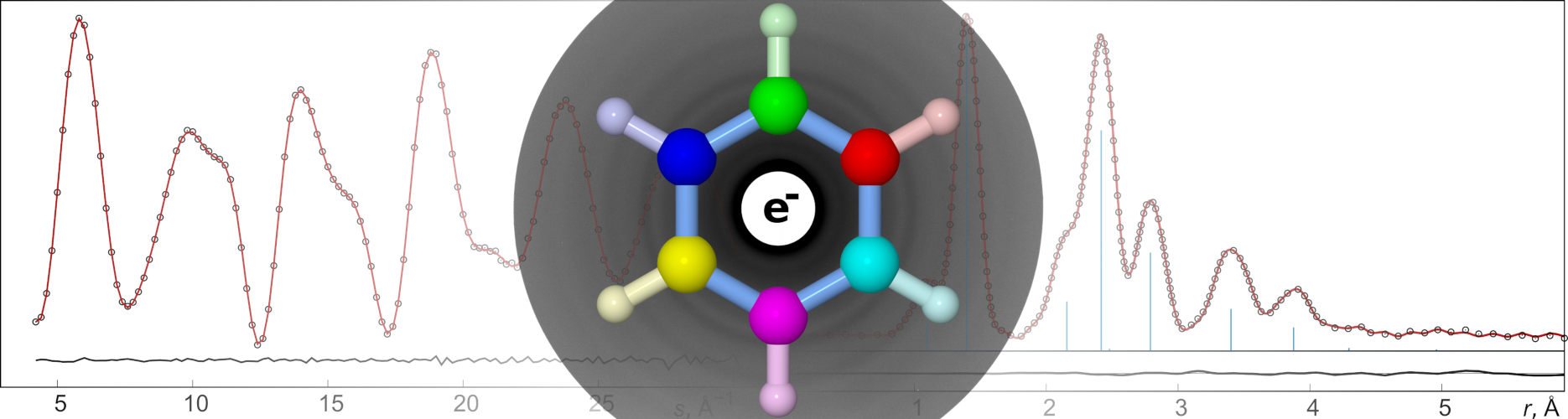
UNEX is a program environment for investigation of molecular structure. It has been created by the Vishnevskiy Group. In this project we develop new and existing experimental methods and combine them to increase accuracy and precision of results. At the current stage the full support for the gas electron diffraction (GED) method is provided, from the calibration of instruments and data reduction to the refinement of molecular structure. Additionally, rotational constants can be used solely or in combination with GED data for determination of molecular geometry. See manual for more details.
Development activity
2024-01-14: Manual updated 2024-01-13: Manual updated 2023-12-13: Added a test for statistical thermodynamics 2023-11-27: Added reference for a MCMIN test 2023-11-25: Adjusted tests 2023-11-25: Hashes for restraining geom. parameters may be calculated 2023-11-25: Added reference files to minimization tests 2023-11-22: Added initial implementation of the msRRHO-2 therm. model 2023-11-18: Implemented quasi-RRHO model for thermodynamic functions 2023-11-17: Fixed test
